Experimenting with Github Actions
 Photo by Luca Bravo on Unsplash
Photo by Luca Bravo on Unsplash
GitHub Actions allow flexible and potentially complicated actions that comprise workflows that respond to repository events on Github. Continuous integration, messaging Slack, greeting new contributors, deploying applications, and many other templates are ready for customization and integration into any repo.
As of this writing, GitHub Actions are available as a beta feature, requiring signup.
Definitions
Action
An action is the smallest portable building block of a workflow. Actions are combined as steps to form a workflow. Actions can be created from scratch, modified from publicly available actions, and shared with others.
Workflow
A workflow is a collection of steps (actions) that describe a process
that will execute in response to GitHub repository events. A YAML
file that defines your workflow configuration with at least one
job. This file lives in the root of your GitHub repository in the
.github/workflows directory. Multiple workflow files may exist and
all will be active.
Step
A step is a set of tasks performed by a job. Each step in a job executes in the same virtual environment, allowing the actions in that job to share information using the filesystem. Steps can run commands (shell commands, etc.) or actions.
Virtual environment
GitHub has hosts for Linux, macOS, and Windows that can serve as execution environments for workflows.
Runner
Jobs run on a service that waits in virtual environment that waits for available jobs. The runner takes jobs one-at-a-time and runs each, reporting logs, etc., back to Github.
Event
Workflows can be triggered by a [repository event] such as a push, a new issue, etc.
Artifact
Artifacts are the files that are created during a workflow. These can be deployed, used in other workflows, published, etc. Additional actions allow working with artifacts.
Walkthrough
Create repo
Here, I simulate creating a simple python package by forking this example repo by bast. The package is to exemplify some best practices for structuring python code. However, here it serves as a simple example of a formal python package that we can build and test.
Login to Github, navigate to the example repo, and fork the repo.
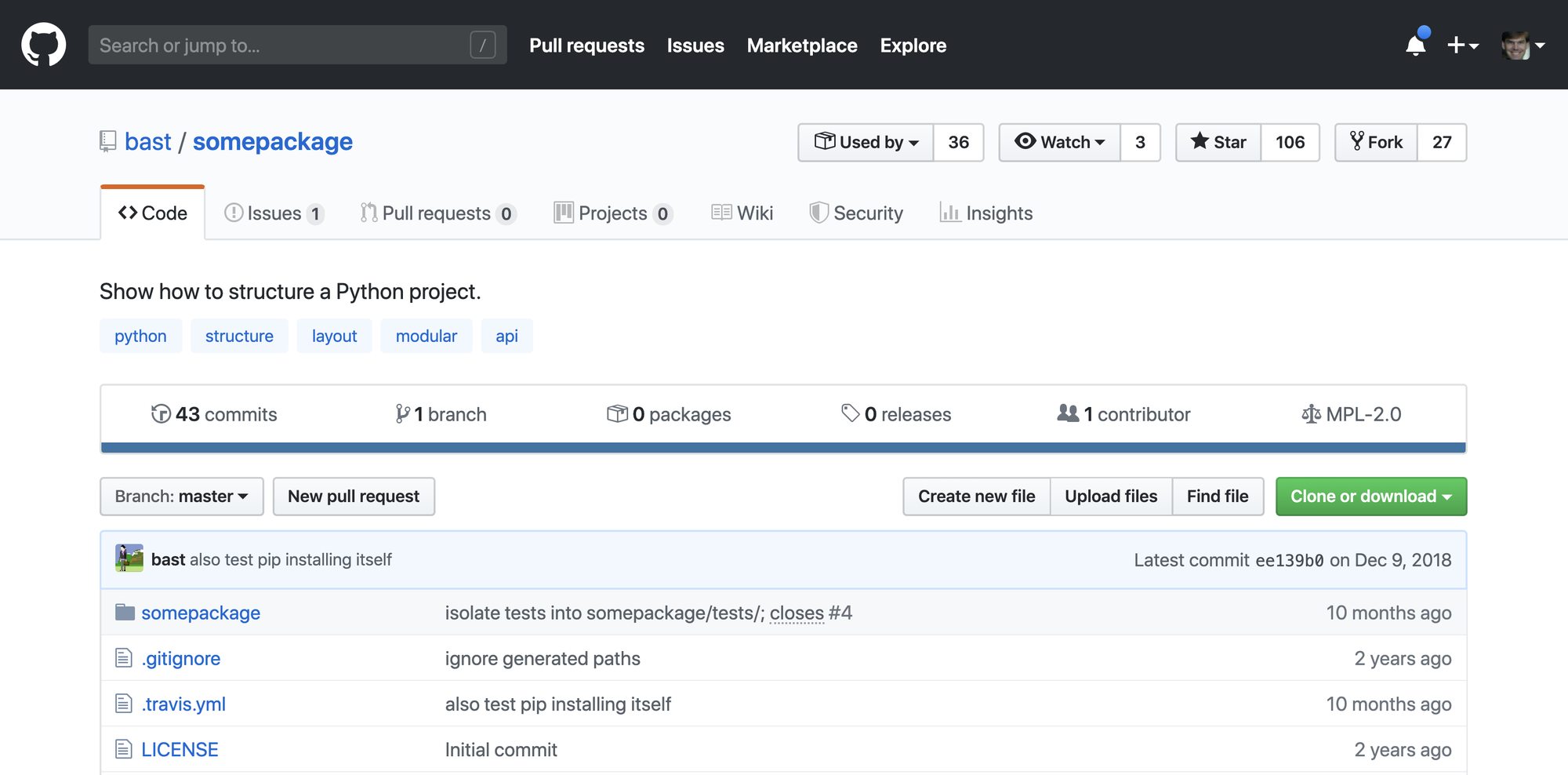
After forking, a fresh package will be available in your repository.
Add a github action
After ensuring that you have [signed up] for Github Actions and have
verified that you are “in” by noticing the Actions button at the top
of your github repository:
Now, we can proceed with adding a workflow from the set of recommended
workflow templates (or others). Clicking on the Actions button will
bring up the following screen.
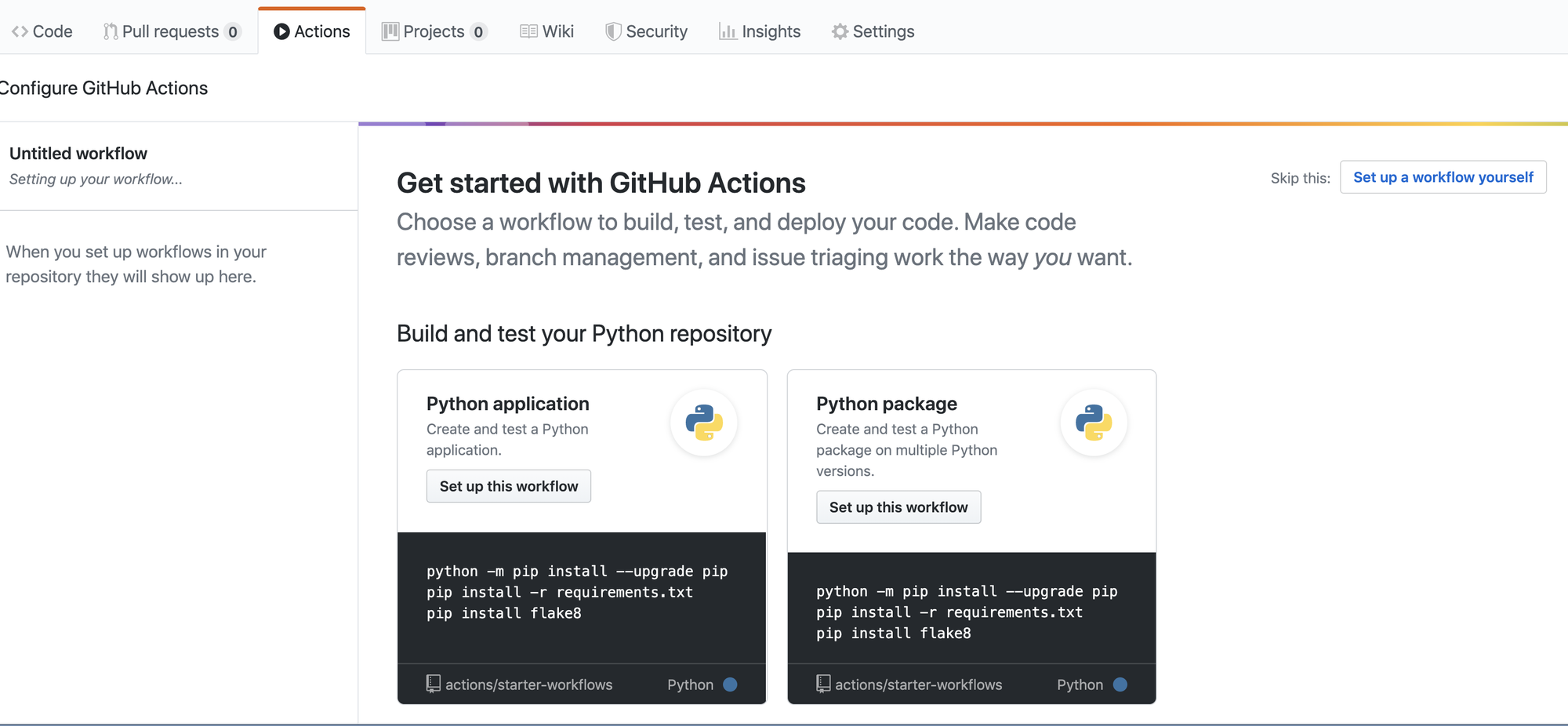
Click on the Set up this workflow on the “Python package”
card. That will bring up the following screen.
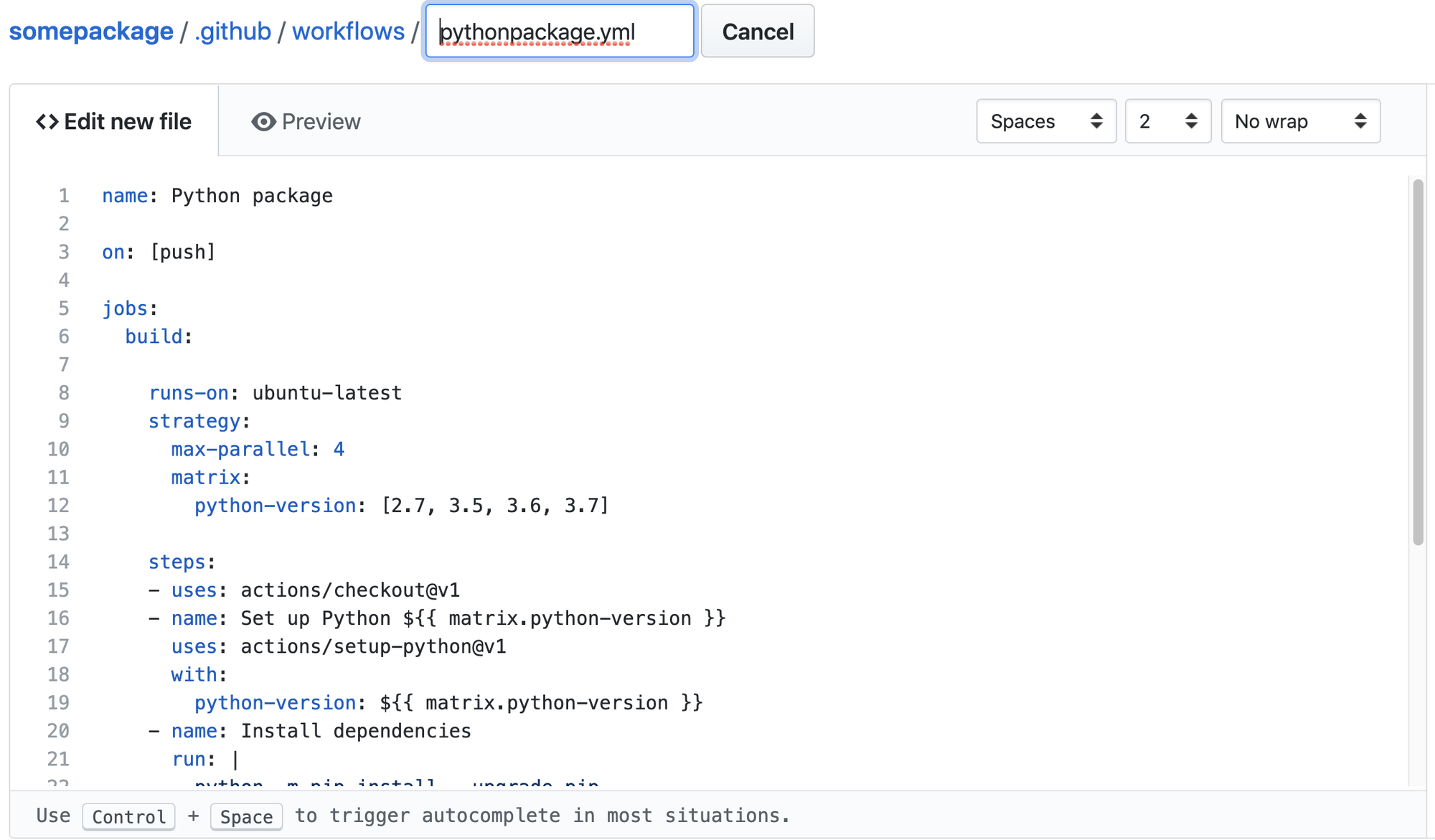
What is happening at this point is that Github is about to add a new
file to your repository. The file will be located at
somepackage/.github/workflows. Edit the filename as you see
fit.
The contents of the workflow file will look like the listing below by default and follows the workflow syntax.
name: Python package
on: [push]
jobs:
build:
runs-on: ubuntu-latest
strategy:
max-parallel: 4
matrix:
python-version: [2.7, 3.5, 3.6, 3.7]
steps:
- uses: actions/checkout@v1
- name: Set up Python ${{ matrix.python-version }}
uses: actions/setup-python@v1
with:
python-version: ${{ matrix.python-version }}
- name: Install dependencies
run: |
python -m pip install --upgrade pip
pip install -r requirements.txt
- name: Lint with flake8
run: |
pip install flake8
# stop the build if there are Python syntax errors or undefined names
flake8 . --count --select=E9,F63,F7,F82 --show-source --statistics
# exit-zero treats all errors as warnings. The GitHub editor is 127 chars wide
flake8 . --count --exit-zero --max-complexity=10 --max-line-length=127 --statistics
- name: Test with pytest
run: |
pip install pytest
pytest
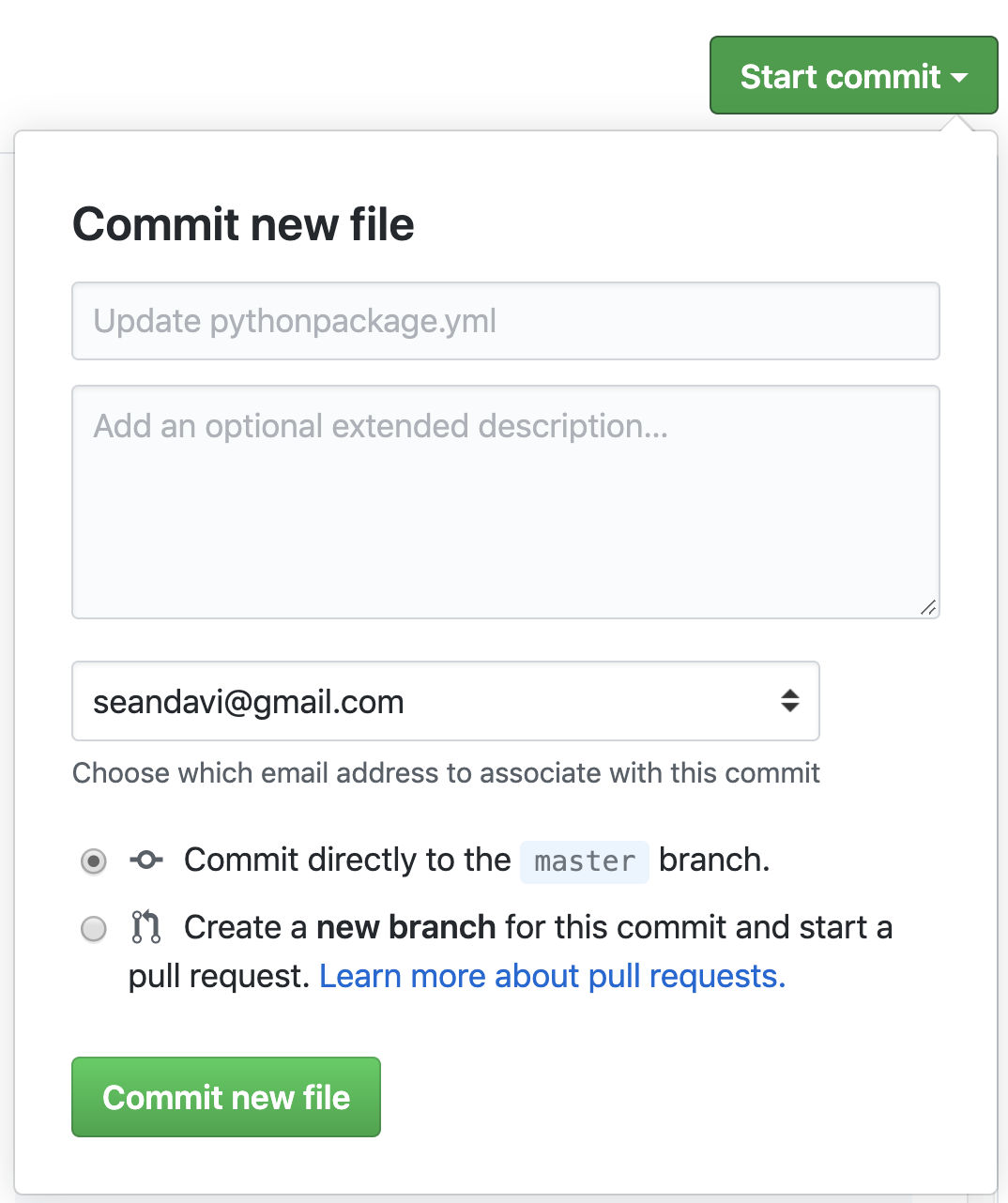
Commit new workflow file
Once you have made any edits (no need to, though), go ahead and click
the Start Commit button.
Build results
By navigating back to Actions, you should see that the workflow is
running or has already completed. An example of what a running
workflow might look like is below.
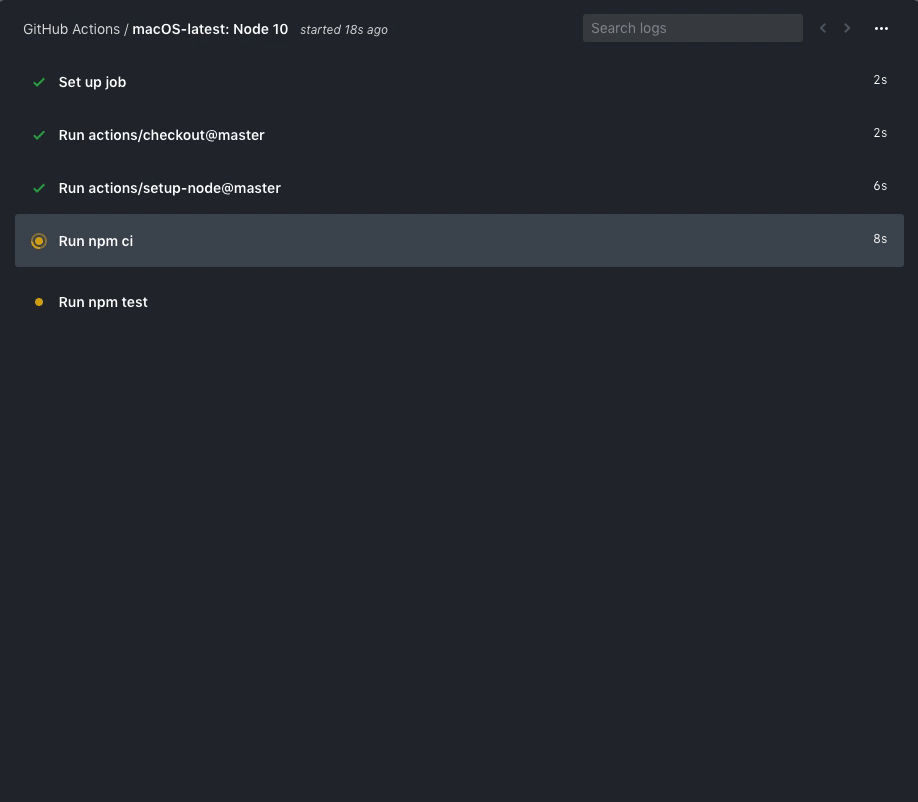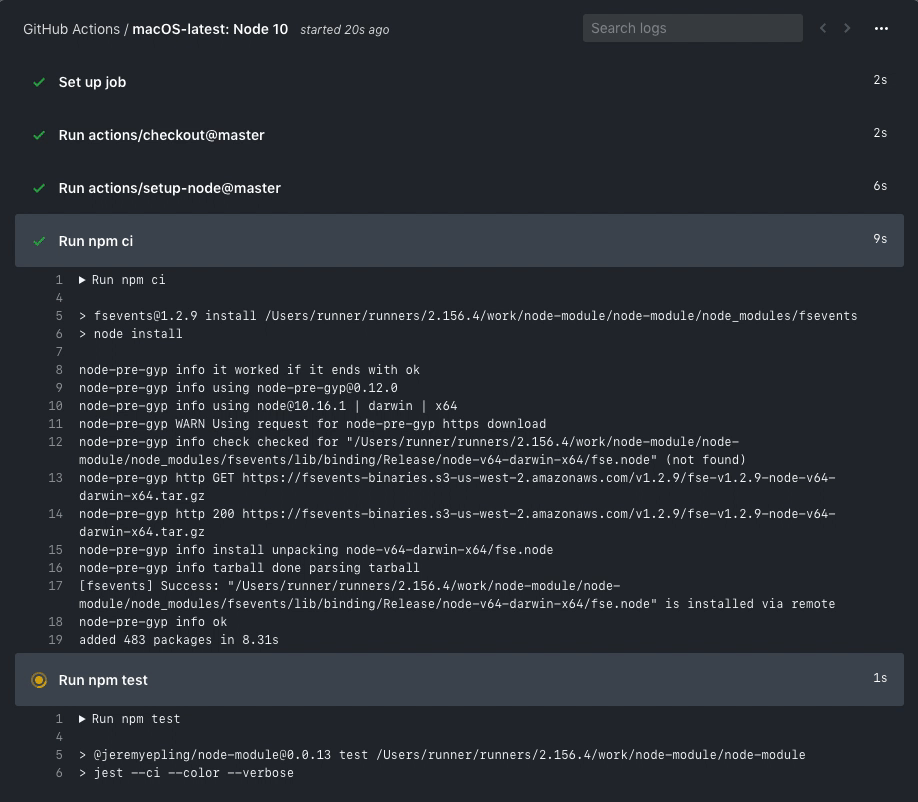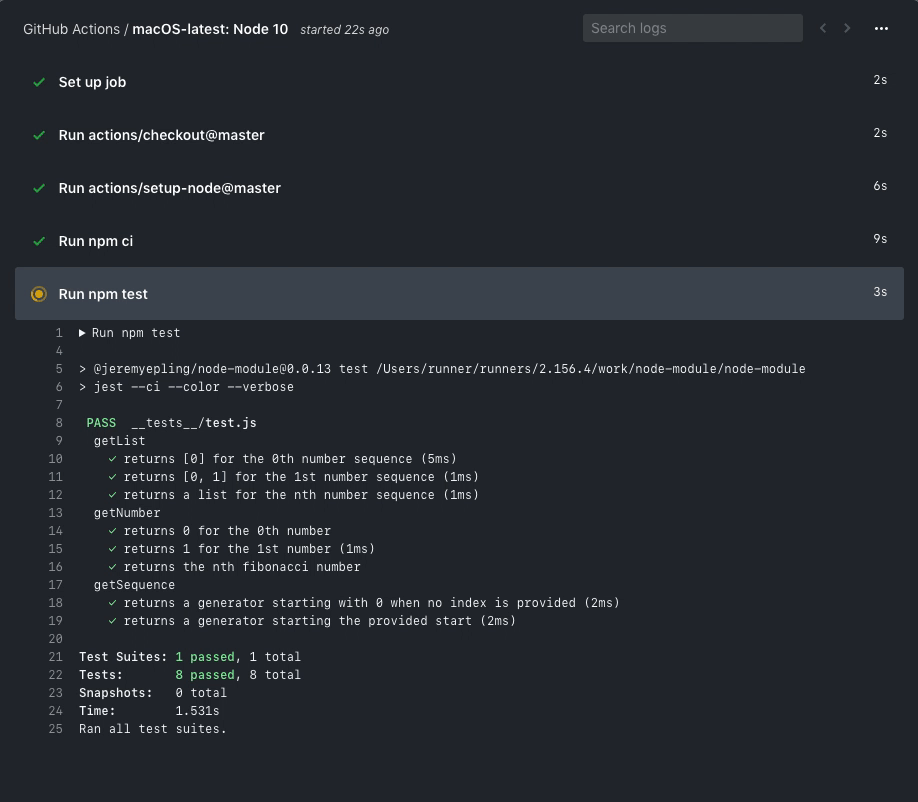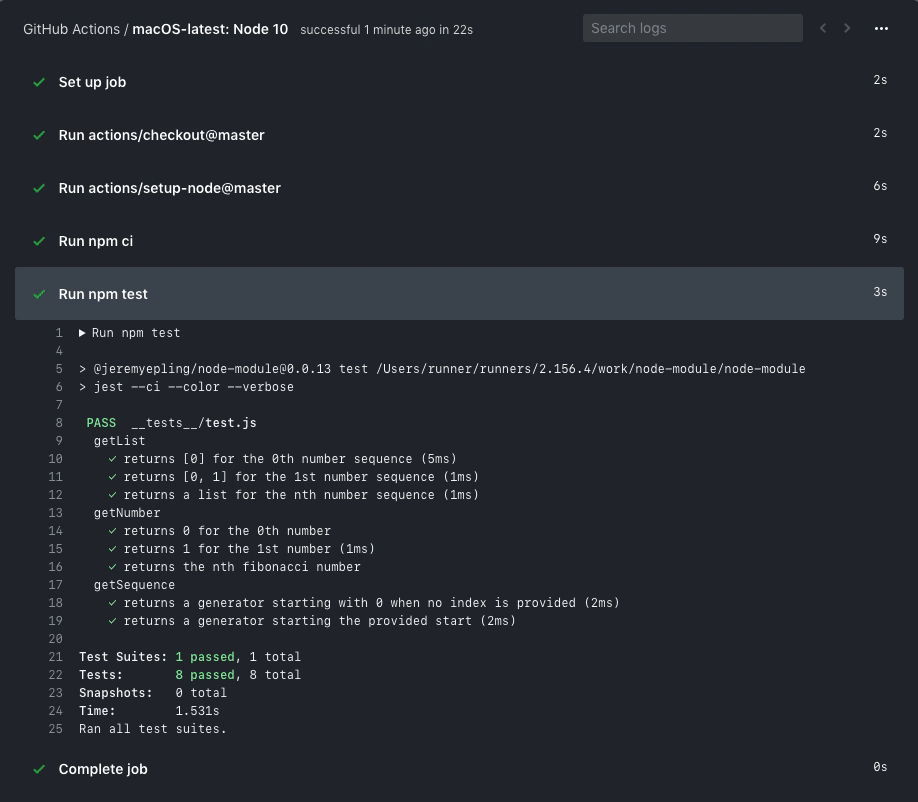
You are free to investigate the logs, etc.
Modifying the workflow
Modifying the workflow is as simple as making edits to the
.github/workflows/...yaml file. For example, you might consider
changing the name of the workflow and modifying the python versions
that are used for testing.
Workflow status badge
Follow these instructions to add a status badge to your repo that will be updated with each workflow execution. Example code is here:
[](https://github.com/seandavi/somepackage/actions)
Note the %20 that represents a url-encoded space character. The resulting badge looks like this:
Conclusion
Github actions and workflows add continuous integration and other capabilities that are configurable via text files that are captured and versioned alongside code. Therefore, actions can be shared, modified, and manipulated with standard text editors, etc.
Many actions are publicly available. Example workflows are also easy to find. For those who are partial to R, there is the R-centric ghactions package that 1) provides workflow templates for common R projects (packages, RMarkdown, …) with sensible defaults and 2) wraps and curates relevant existing external actions, such as those to deploy to GitHub Pages or Netlify.
Hardware available as of this writing is described here and includes Windows, Linux, and MacOS environments. When a workflow is running, multiple environment variables are set and accessible to software. Github actions allow the user to create secrets to securely store credentials for use inside run environments. Github actions are container-ready, allowing containerized actions.
This post, like many of my posts, is very operational, but I have found that a worked example is usually more valuable than long expository posts. That said, I always appreciate comments and suggestions for improvements (see below).